HAB Observing Network – New England
The HABON-NE (HAB Observing Network - New England) program is a framework for the deployment of a fleet of advanced sensors and sensor platforms in New England waters. The goals of this project are to improve HAB surveillance and ,support resource management decision-making. HABON-NE sensor deployments are year-round, at key locations across the region (see Fig. 1, right). In addition to these static deployments, novel mobile platforms will enable distribution of sensor assets to address seasonal shifts in HAB geography, as well as rapid redeployment when blooms develop in unexpected areas. Adaptability is an important part of this program, and autonomous sensors can be relocated or sampling frequency adapted in response to changing seasonal as well as unexpected new ones.
The network’s core sensors are two advanced robotic instruments, the Imaging FlowCytobot (IFCB) and the Environmental Sample Processor (ESP). Both discriminate HAB cells from non-harmful phytoplankton but differ in their approaches to detection and quantification. Inherent in these differences are complementary capabilities and trade-offs between species-specificity, observing frequency, power consumption, and cost of operation. The project aims to leverage the sensors’ complementary aspects to best provide timely, cost-effective, and actionable information to stakeholders.
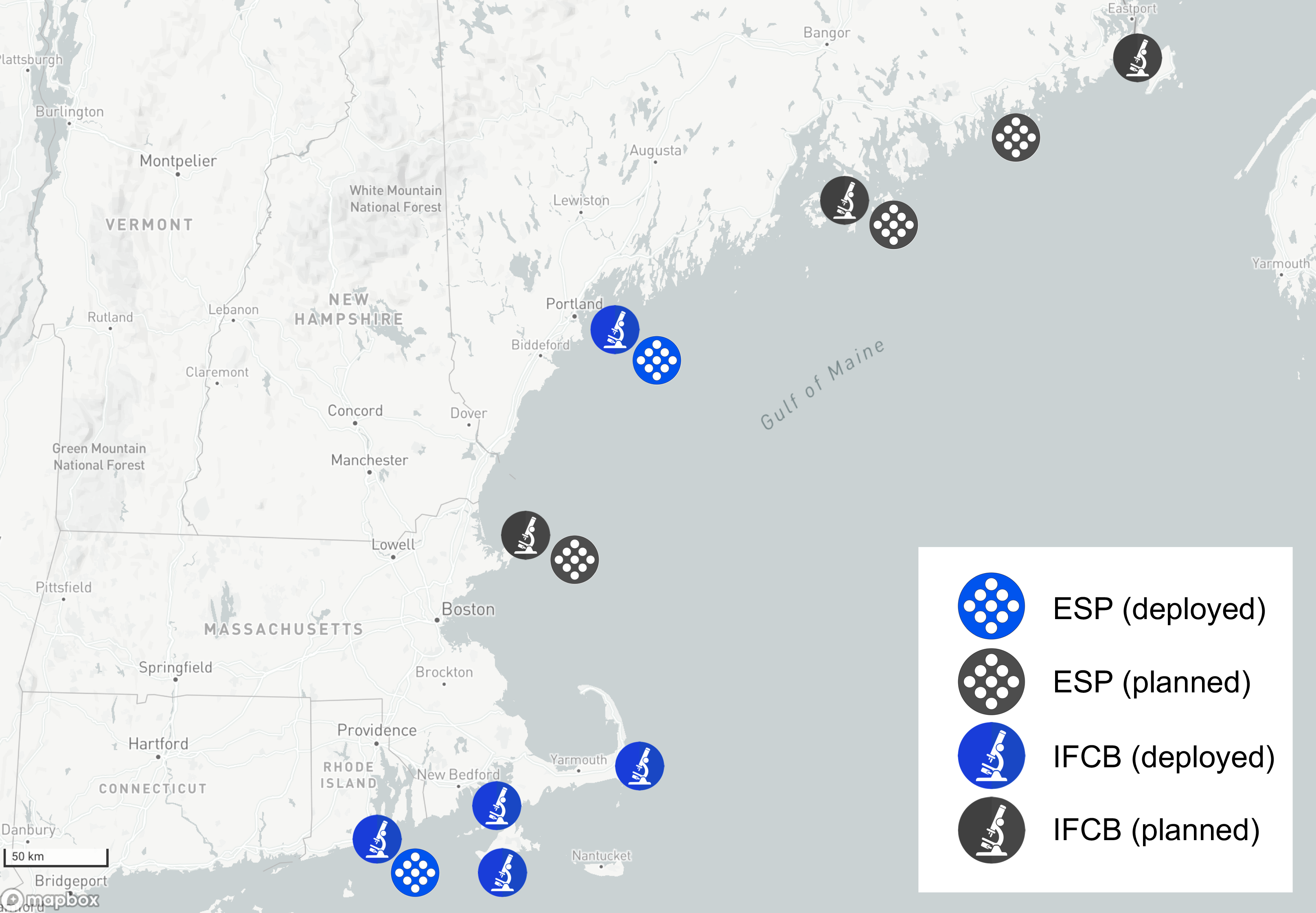
Locations of current and planned Imaging FlowCytobot (IFCB) and Environmental Sample Processor (ESP) sensor installations in New England. The HABON-NE sensor assess include five IFCBs, six ESPs including 1 spESP variant, two trailered PhytO-ARM platforms, and an SP-48 SeaTrac boat for mobile deployments.
Observations will be shared as they are produced through the WHOI HAB Hub data portal, thus improving situational awareness throughout the region. The WHOI HAB Hub is being developed as an open source, containerized webserver platform for region-scale integration and sharing of rich and diversely sourced HAB observations, model outputs, contextual data, and management actions. Going forward, the project will also implement and extend a toxicity model that translates high frequency, in situ estimates of cell concentrations into estimates of PSP toxin loads in shellfish.
